
A | B | C | D | E | F | G | H | CH | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9
| Part of a series on |
| Genetics |
|---|
 |
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery.[1][2]
Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers,[3] characterize antibody repertoire,[4] and can be used to guide patient treatment.[5] Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged.[4]
The rapid speed of sequencing attained with modern DNA sequencing technology has been instrumental in the sequencing of complete DNA sequences, or genomes, of numerous types and species of life, including the human genome and other complete DNA sequences of many animal, plant, and microbial species.
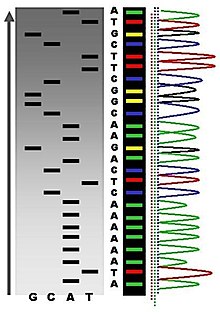
The first DNA sequences were obtained in the early 1970s by academic researchers using laborious methods based on two-dimensional chromatography. Following the development of fluorescence-based sequencing methods with a DNA sequencer,[6] DNA sequencing has become easier and orders of magnitude faster.[7][8]
Applications
DNA sequencing may be used to determine the sequence of individual genes, larger genetic regions (i.e. clusters of genes or operons), full chromosomes, or entire genomes of any organism. DNA sequencing is also the most efficient way to indirectly sequence RNA or proteins (via their open reading frames). In fact, DNA sequencing has become a key technology in many areas of biology and other sciences such as medicine, forensics, and anthropology.
Molecular biology
Sequencing is used in molecular biology to study genomes and the proteins they encode. Information obtained using sequencing allows researchers to identify changes in genes and noncoding DNA (including regulatory sequences), associations with diseases and phenotypes, and identify potential drug targets.
Evolutionary biology
Since DNA is an informative macromolecule in terms of transmission from one generation to another, DNA sequencing is used in evolutionary biology to study how different organisms are related and how they evolved. In February 2021, scientists reported, for the first time, the sequencing of DNA from animal remains, a mammoth in this instance, over a million years old, the oldest DNA sequenced to date.[9][10]
Metagenomics
The field of metagenomics involves identification of organisms present in a body of water, sewage, dirt, debris filtered from the air, or swab samples from organisms. Knowing which organisms are present in a particular environment is critical to research in ecology, epidemiology, microbiology, and other fields. Sequencing enables researchers to determine which types of microbes may be present in a microbiome, for example.
Virology
As most viruses are too small to be seen by a light microscope, sequencing is one of the main tools in virology to identify and study the virus.[11] Viral genomes can be based in DNA or RNA. RNA viruses are more time-sensitive for genome sequencing, as they degrade faster in clinical samples.[12] Traditional Sanger sequencing and next-generation sequencing are used to sequence viruses in basic and clinical research, as well as for the diagnosis of emerging viral infections, molecular epidemiology of viral pathogens, and drug-resistance testing. There are more than 2.3 million unique viral sequences in GenBank.[11] Recently, NGS has surpassed traditional Sanger as the most popular approach for generating viral genomes.[11]
During the 1990 avian influenza outbreak, viral sequencing determined that the influenza sub-type originated through reassortment between quail and poultry. This led to legislation in Hong Kong that prohibited selling live quail and poultry together at market. Viral sequencing can also be used to estimate when a viral outbreak began by using a molecular clock technique.[12]
Medicine
Medical technicians may sequence genes (or, theoretically, full genomes) from patients to determine if there is risk of genetic diseases. This is a form of genetic testing, though some genetic tests may not involve DNA sequencing.
As of 2013 DNA sequencing was increasingly used to diagnose and treat rare diseases. As more and more genes are identified that cause rare genetic diseases, molecular diagnoses for patients become more mainstream. DNA sequencing allows clinicians to identify genetic diseases, improve disease management, provide reproductive counseling, and more effective therapies.[13] Gene sequencing panels are used to identify multiple potential genetic causes of a suspected disorder.[14]
Also, DNA sequencing may be useful for determining a specific bacteria, to allow for more precise antibiotics treatments, hereby reducing the risk of creating antimicrobial resistance in bacteria populations.[15][16][17][18][19][20]
Forensic investigation
DNA sequencing may be used along with DNA profiling methods for forensic identification[21] and paternity testing. DNA testing has evolved tremendously in the last few decades to ultimately link a DNA print to what is under investigation. The DNA patterns in fingerprint, saliva, hair follicles, etc. uniquely separate each living organism from another. Testing DNA is a technique which can detect specific genomes in a DNA strand to produce a unique and individualized pattern.
The four canonical bases
The canonical structure of DNA has four bases: thymine (T), adenine (A), cytosine (C), and guanine (G). DNA sequencing is the determination of the physical order of these bases in a molecule of DNA. However, there are many other bases that may be present in a molecule. In some viruses (specifically, bacteriophage), cytosine may be replaced by hydroxy methyl or hydroxy methyl glucose cytosine.[22] In mammalian DNA, variant bases with methyl groups or phosphosulfate may be found.[23][24] Depending on the sequencing technique, a particular modification, e.g., the 5mC (5 methyl cytosine) common in humans, may or may not be detected.[25]
In almost all organisms, DNA is synthesized in vivo using only the 4 canonical bases; modification that occurs post replication creates other bases like 5 methyl C. However, some bacteriophage can incorporate a non standard base directly.[26]
In addittion to modifications, DNA is under constant assault by environmental agents such as UV and Oxygen radicals. At the present time, the presence of such damaged bases is not detected by most DNA sequencing methods, although PacBio has published on this https://www.pacb.com/publications/direct-detection-and-sequencing-of-damaged-dna-bases/
History
Discovery of DNA structure and function
Deoxyribonucleic acid (DNA) was first discovered and isolated by Friedrich Miescher in 1869, but it remained under-studied for many decades because proteins, rather than DNA, were thought to hold the genetic blueprint to life. This situation changed after 1944 as a result of some experiments by Oswald Avery, Colin MacLeod, and Maclyn McCarty demonstrating that purified DNA could change one strain of bacteria into another. This was the first time that DNA was shown capable of transforming the properties of cells.
In 1953, James Watson and Francis Crick put forward their double-helix model of DNA, based on crystallized X-ray structures being studied by Rosalind Franklin. According to the model, DNA is composed of two strands of nucleotides coiled around each other, linked together by hydrogen bonds and running in opposite directions. Each strand is composed of four complementary nucleotides – adenine (A), cytosine (C), guanine (G) and thymine (T) – with an A on one strand always paired with T on the other, and C always paired with G. They proposed that such a structure allowed each strand to be used to reconstruct the other, an idea central to the passing on of hereditary information between generations.[27]

The foundation for sequencing proteins was first laid by the work of Frederick Sanger who by 1955 had completed the sequence of all the amino acids in insulin, a small protein secreted by the pancreas. This provided the first conclusive evidence that proteins were chemical entities with a specific molecular pattern rather than a random mixture of material suspended in fluid. Sanger's success in sequencing insulin spurred on x-ray crystallographers, including Watson and Crick, who by now were trying to understand how DNA directed the formation of proteins within a cell. Soon after attending a series of lectures given by Frederick Sanger in October 1954, Crick began developing a theory which argued that the arrangement of nucleotides in DNA determined the sequence of amino acids in proteins, which in turn helped determine the function of a protein. He published this theory in 1958.[28]
RNA sequencing
RNA sequencing was one of the earliest forms of nucleotide sequencing. The major landmark of RNA sequencing is the sequence of the first complete gene and the complete genome of Bacteriophage MS2, identified and published by Walter Fiers and his coworkers at the University of Ghent (Ghent, Belgium), in 1972[29] and 1976.[30] Traditional RNA sequencing methods require the creation of a cDNA molecule which must be sequenced.[31]
Early DNA sequencing methods
The first method for determining DNA sequences involved a location-specific primer extension strategy established by Ray Wu at Cornell University in 1970.[32] DNA polymerase catalysis and specific nucleotide labeling, both of which figure prominently in current sequencing schemes, were used to sequence the cohesive ends of lambda phage DNA.[33][34][35] Between 1970 and 1973, Wu, R Padmanabhan and colleagues demonstrated that this method can be employed to determine any DNA sequence using synthetic location-specific primers.[36][37][8] Frederick Sanger then adopted this primer-extension strategy to develop more rapid DNA sequencing methods at the MRC Centre, Cambridge, UK and published a method for "DNA sequencing with chain-terminating inhibitors" in 1977.[38] Walter Gilbert and Allan Maxam at Harvard also developed sequencing methods, including one for "DNA sequencing by chemical degradation".[39][40] In 1973, Gilbert and Maxam reported the sequence of 24 basepairs using a method known as wandering-spot analysis.[41] Advancements in sequencing were aided by the concurrent development of recombinant DNA technology, allowing DNA samples to be isolated from sources other than viruses.
Sequencing of full genomes

The first full DNA genome to be sequenced was that of bacteriophage φX174 in 1977.[42] Medical Research Council scientists deciphered the complete DNA sequence of the Epstein-Barr virus in 1984, finding it contained 172,282 nucleotides. Completion of the sequence marked a significant turning point in DNA sequencing because it was achieved with no prior genetic profile knowledge of the virus.[43][8]
A non-radioactive method for transferring the DNA molecules of sequencing reaction mixtures onto an immobilizing matrix during electrophoresis was developed by Herbert Pohl and co-workers in the early 1980s.[44][45] Followed by the commercialization of the DNA sequencer "Direct-Blotting-Electrophoresis-System GATC 1500" by GATC Biotech, which was intensively used in the framework of the EU genome-sequencing programme, the complete DNA sequence of the yeast Saccharomyces cerevisiae chromosome II.[46] Leroy E. Hood's laboratory at the California Institute of Technology announced the first semi-automated DNA sequencing machine in 1986.[47] This was followed by Applied Biosystems' marketing of the first fully automated sequencing machine, the ABI 370, in 1987 and by Dupont's Genesis 2000[48] which used a novel fluorescent labeling technique enabling all four dideoxynucleotides to be identified in a single lane. By 1990, the U.S. National Institutes of Health (NIH) had begun large-scale sequencing trials on Mycoplasma capricolum, Escherichia coli, Caenorhabditis elegans, and Saccharomyces cerevisiae at a cost of US$0.75 per base. Meanwhile, sequencing of human cDNA sequences called expressed sequence tags began in Craig Venter's lab, an attempt to capture the coding fraction of the human genome.[49] In 1995, Venter, Hamilton Smith, and colleagues at The Institute for Genomic Research (TIGR) published the first complete genome of a free-living organism, the bacterium Haemophilus influenzae. The circular chromosome contains 1,830,137 bases and its publication in the journal Science[50] marked the first published use of whole-genome shotgun sequencing, eliminating the need for initial mapping efforts.
By 2001, shotgun sequencing methods had been used to produce a draft sequence of the human genome.[51][52]
High-throughput sequencing (HTS) methods

Several new methods for DNA sequencing were developed in the mid to late 1990s and were implemented in commercial DNA sequencers by 2000. Together these were called the "next-generation" or "second-generation" sequencing (NGS) methods, in order to distinguish them from the earlier methods, including Sanger sequencing. In contrast to the first generation of sequencing, NGS technology is typically characterized by being highly scalable, allowing the entire genome to be sequenced at once. Usually, this is accomplished by fragmenting the genome into small pieces, randomly sampling for a fragment, and sequencing it using one of a variety of technologies, such as those described below. An entire genome is possible because multiple fragments are sequenced at once (giving it the name "massively parallel" sequencing) in an automated process.
NGS technology has tremendously empowered researchers to look for insights into health, anthropologists to investigate human origins, and is catalyzing the "Personalized Medicine" movement. However, it has also opened the door to more room for error. There are many software tools to carry out the computational analysis of NGS data, often compiled at online platforms such as CSI NGS Portal, each with its own algorithm. Even the parameters within one software package can change the outcome of the analysis. In addition, the large quantities of data produced by DNA sequencing have also required development of new methods and programs for sequence analysis. Several efforts to develop standards in the NGS field have been attempted to address these challenges, most of which have been small-scale efforts arising from individual labs. Most recently, a large, organized, FDA-funded effort has culminated in the BioCompute standard.
On 26 October 1990, Roger Tsien, Pepi Ross, Margaret Fahnestock and Allan J Johnston filed a patent describing stepwise ("base-by-base") sequencing with removable 3' blockers on DNA arrays (blots and single DNA molecules).[54] In 1996, Pål Nyrén and his student Mostafa Ronaghi at the Royal Institute of Technology in Stockholm published their method of pyrosequencing.[55]
On 1 April 1997, Pascal Mayer and Laurent Farinelli submitted patents to the World Intellectual Property Organization describing DNA colony sequencing.[56] The DNA sample preparation and random surface-polymerase chain reaction (PCR) arraying methods described in this patent, coupled to Roger Tsien et al.'s "base-by-base" sequencing method, is now implemented in Illumina's Hi-Seq genome sequencers.
In 1998, Phil Green and Brent Ewing of the University of Washington described their phred quality score for sequencer data analysis,[57] a landmark analysis technique that gained widespread adoption, and which is still the most common metric for assessing the accuracy of a sequencing platform.[58]
Lynx Therapeutics published and marketed massively parallel signature sequencing (MPSS), in 2000. This method incorporated a parallelized, adapter/ligation-mediated, bead-based sequencing technology and served as the first commercially available "next-generation" sequencing method, though no DNA sequencers were sold to independent laboratories.[59]
Basic methods
Maxam-Gilbert sequencing
Allan Maxam and Walter Gilbert published a DNA sequencing method in 1977 based on chemical modification of DNA and subsequent cleavage at specific bases.[39] Also known as chemical sequencing, this method allowed purified samples of double-stranded DNA to be used without further cloning. This method's use of radioactive labeling and its technical complexity discouraged extensive use after refinements in the Sanger methods had been made.
Maxam-Gilbert sequencing requires radioactive labeling at one 5' end of the DNA and purification of the DNA fragment to be sequenced. Chemical treatment then generates breaks at a small proportion of one or two of the four nucleotide bases in each of four reactions (G, A+G, C, C+T). The concentration of the modifying chemicals is controlled to introduce on average one modification per DNA molecule. Thus a series of labeled fragments is generated, from the radiolabeled end to the first "cut" site in each molecule. The fragments in the four reactions are electrophoresed side by side in denaturing acrylamide gels for size separation. To visualize the fragments, the gel is exposed to X-ray film for autoradiography, yielding a series of dark bands each corresponding to a radiolabeled DNA fragment, from which the sequence may be inferred.[39]
This method is mostly obsolete as of 2023.[60]
Chain-termination methods
The chain-termination method developed by Frederick Sanger and coworkers in 1977 soon became the method of choice, owing to its relative ease and reliability.[38][61] When invented, the chain-terminator method used fewer toxic chemicals and lower amounts of radioactivity than the Maxam and Gilbert method. Because of its comparative ease, the Sanger method was soon automated and was the method used in the first generation of DNA sequencers.
Sanger sequencing is the method which prevailed from the 1980s until the mid-2000s. Over that period, great advances were made in the technique, such as fluorescent labelling, capillary electrophoresis, and general automation. These developments allowed much more efficient sequencing, leading to lower costs. The Sanger method, in mass production form, is the technology which produced the first human genome in 2001, ushering in the age of genomics. However, later in the decade, radically different approaches reached the market, bringing the cost per genome down from $100 million in 2001 to $10,000 in 2011.[62]
Sequencing by synthesis
The objective for sequential sequencing by synthesis (SBS) is to determine the sequencing of a DNA sample by detecting the incorporation of a nucleotide by a DNA polymerase. An engineered polymerase is used to synthesize a copy of a single strand of DNA and the incorporation of each nucleotide is monitored. The principle of real-time sequencing by synthesis was first described in 1993[63] with improvements published some years later.[64] The key parts are highly similar for all embodiments of SBS and includes (1) amplification of DNA (to enhance the subsequent signal) and attach the DNA to be sequenced to a solid support, (2) generation of single stranded DNA on the solid support, (3) incorporation of nucleotides using an engineered polymerase and (4) real-time detection of the incorporation of nucleotide The steps 3-4 are repeated and the sequence is assembled from the signals obtained in step 4. This principle of real-time sequencing-by-synthesis has been used for almost all massive parallel sequencing instruments, including 454, PacBio, IonTorrent, Illumina and MGI.
Large-scale sequencing and de novo sequencing
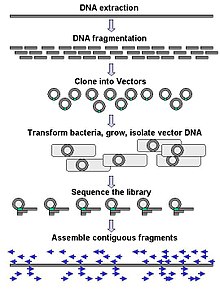
Large-scale sequencing often aims at sequencing very long DNA pieces, such as whole chromosomes, although large-scale sequencing can also be used to generate very large numbers of short sequences, such as found in phage display. For longer targets such as chromosomes, common approaches consist of cutting (with restriction enzymes) or shearing (with mechanical forces) large DNA fragments into shorter DNA fragments. The fragmented DNA may then be cloned into a DNA vector and amplified in a bacterial host such as Escherichia coli. Short DNA fragments purified from individual bacterial colonies are individually sequenced and assembled electronically into one long, contiguous sequence. Studies have shown that adding a size selection step to collect DNA fragments of uniform size can improve sequencing efficiency and accuracy of the genome assembly. In these studies, automated sizing has proven to be more reproducible and precise than manual gel sizing.[65][66][67]
The term "de novo sequencing" specifically refers to methods used to determine the sequence of DNA with no previously known sequence. De novo translates from Latin as "from the beginning". Gaps in the assembled sequence may be filled by primer walking. The different strategies have different tradeoffs in speed and accuracy; shotgun methods are often used for sequencing large genomes, but its assembly is complex and difficult, particularly with sequence repeats often causing gaps in genome assembly.
Most sequencing approaches use an in vitro cloning step to amplify individual DNA molecules, because their molecular detection methods are not sensitive enough for single molecule sequencing. Emulsion PCR[68] isolates individual DNA molecules along with primer-coated beads in aqueous droplets within an oil phase. A polymerase chain reaction (PCR) then coats each bead with clonal copies of the DNA molecule followed by immobilization for later sequencing. Emulsion PCR is used in the methods developed by Marguilis et al. (commercialized by 454 Life Sciences), Shendure and Porreca et al. (also known as "polony sequencing") and SOLiD sequencing, (developed by Agencourt, later Applied Biosystems, now Life Technologies).[69][70][71] Emulsion PCR is also used in the GemCode and Chromium platforms developed by 10x Genomics.[72]
Shotgun sequencing
Shotgun sequencing is a sequencing method designed for analysis of DNA sequences longer than 1000 base pairs, up to and including entire chromosomes. This method requires the target DNA to be broken into random fragments. After sequencing individual fragments using the chain termination method, the sequences can be reassembled on the basis of their overlapping regions.[73]
High-throughput methods

High-throughput sequencing, which includes next-generation "short-read" and third-generation "long-read" sequencing methods,[nt 1] applies to exome sequencing, genome sequencing, genome resequencing, transcriptome profiling (RNA-Seq), DNA-protein interactions (ChIP-sequencing), and epigenome characterization.[74]
The high demand for low-cost sequencing has driven the development of high-throughput sequencing technologies that parallelize the sequencing process, producing thousands or millions of sequences concurrently.[75][76][77] High-throughput sequencing technologies are intended to lower the cost of DNA sequencing beyond what is possible with standard dye-terminator methods.[78] In ultra-high-throughput sequencing as many as 500,000 sequencing-by-synthesis operations may be run in parallel.[79][80][81] Such technologies led to the ability to sequence an entire human genome in as little as one day.[82] As of 2019[update], corporate leaders in the development of high-throughput sequencing products included Illumina, Qiagen and ThermoFisher Scientific.[82]
| Method | Read length | Accuracy (single read not consensus) | Reads per run | Time per run | Cost per 1 billion bases (in US$) | Advantages | Disadvantages |
|---|---|---|---|---|---|---|---|
| Single-molecule real-time sequencing (Pacific Biosciences) | 30,000 bp (N50); | 87% raw-read accuracy[88] | 4,000,000 per Sequel 2 SMRT cell, 100–200 gigabases[85][89][90] | 30 minutes to 20 hours[85][91] | $7.2-$43.3 | Fast. Detects 4mC, 5mC, 6mA.[92] | Moderate throughput. Equipment can be very expensive. |
| Ion semiconductor (Ion Torrent sequencing) | up to 600 bp[93] | 99.6%[94] | up to 80 million | 2 hours | $66.8-$950 | Less expensive equipment. Fast. | Homopolymer errors. |
| Pyrosequencing (454) | 700 bp | 99.9% | 1 million | 24 hours | $10,000 | Long read size. Fast. | Runs are expensive. Homopolymer errors. |
| Sequencing by synthesis (Illumina) | MiniSeq, NextSeq: 75–300 bp;
MiSeq: 50–600 bp; HiSeq 2500: 50–500 bp; HiSeq 3/4000: 50–300 bp; HiSeq X: 300 bp |
99.9% (Phred30) | MiniSeq/MiSeq: 1–25 Million;
NextSeq: 130-00 Million; HiSeq 2500: 300 million – 2 billion; HiSeq 3/4000 2.5 billion; HiSeq X: 3 billion |
1 to 11 days, depending upon sequencer and specified read length[95] | $5 to $150 | Potential for high sequence yield, depending upon sequencer model and desired application. | Equipment can be very expensive. Requires high concentrations of DNA. |
| Combinatorial probe anchor synthesis (cPAS- BGI/MGI) | BGISEQ-50: 35-50bp;
MGISEQ 200: 50-200bp; BGISEQ-500, MGISEQ-2000: 50-300bp[96] |
99.9% (Phred30) | BGISEQ-50: 160M;
MGISEQ 200: 300M; BGISEQ-500: 1300M per flow cell; MGISEQ-2000: 375M FCS flow cell, 1500M FCL flow cell per flow cell. |
1 to 9 days depending on instrument, read length and number of flow cells run at a time. | $5– $120 | ||
| Sequencing by ligation (SOLiD sequencing) | 50+35 or 50+50 bp | 99.9% | 1.2 to 1.4 billion | 1 to 2 weeks | $60–130 | Low cost per base. | Slower than other methods. Has issues sequencing palindromic sequences.[97] |
| Nanopore Sequencing | Dependent on library preparation, not the device, so user chooses read length (up to 2,272,580 bp reported[98]). | ~92–97% single read | dependent on read length selected by user | data streamed in real time. Choose 1 min to 48 hrs | $7–100 | Longest individual reads. Accessible user community. Portable (Palm sized). | Lower throughput than other machines, Single read accuracy in 90s. |
| GenapSys Sequencing | Around 150 bp single-end | 99.9% (Phred30) | 1 to 16 million | Around 24 hours | $667 | Low-cost of instrument ($10,000) | |
| Chain termination (Sanger sequencing) | 400 to 900 bp | 99.9% | N/A | 20 minutes to 3 hours | $2,400,000 | Useful for many applications. | More expensive and impractical for larger sequencing projects. This method also requires the time-consuming step of plasmid cloning or PCR. |
Long-read sequencing methods
Single molecule real time (SMRT) sequencing
SMRT sequencing is based on the sequencing by synthesis approach. The DNA is synthesized in zero-mode wave-guides (ZMWs) – small well-like containers with the capturing tools located at the bottom of the well. The sequencing is performed with use of unmodified polymerase (attached to the ZMW bottom) and fluorescently labelled nucleotides flowing freely in the solution. The wells are constructed in a way that only the fluorescence occurring by the bottom of the well is detected. The fluorescent label is detached from the nucleotide upon its incorporation into the DNA strand, leaving an unmodified DNA strand. According to Pacific Biosciences (PacBio), the SMRT technology developer, this methodology allows detection of nucleotide modifications (such as cytosine methylation). This happens through the observation of polymerase kinetics. This approach allows reads of 20,000 nucleotides or more, with average read lengths of 5 kilobases.[89][99] In 2015, Pacific Biosciences announced the launch of a new sequencing instrument called the Sequel System, with 1 million ZMWs compared to 150,000 ZMWs in the PacBio RS II instrument.[100][101] SMRT sequencing is referred to as "third-generation" or "long-read" sequencing.
Nanopore DNA sequencingedit
The DNA passing through the nanopore changes its ion current. This change is dependent on the shape, size and length of the DNA sequence. Each type of the nucleotide blocks the ion flow through the pore for a different period of time. The method does not require modified nucleotides and is performed in real time. Nanopore sequencing is referred to as "third-generation" or "long-read" sequencing, along with SMRT sequencing.
Early industrial research into this method was based on a technique called 'exonuclease sequencing', where the readout of electrical signals occurred as nucleotides passed by alpha(α)-hemolysin pores covalently bound with cyclodextrin.[102] However the subsequent commercial method, 'strand sequencing', sequenced DNA bases in an intact strand.
Two main areas of nanopore sequencing in development are solid state nanopore sequencing, and protein based nanopore sequencing. Protein nanopore sequencing utilizes membrane protein complexes such as α-hemolysin, MspA (Mycobacterium smegmatis Porin A) or CssG, which show great promise given their ability to distinguish between individual and groups of nucleotides.[103] In contrast, solid-state nanopore sequencing utilizes synthetic materials such as silicon nitride and aluminum oxide and it is preferred for its superior mechanical ability and thermal and chemical stability.[104] The fabrication method is essential for this type of sequencing given that the nanopore array can contain hundreds of pores with diameters smaller than eight nanometers.[103]
The concept originated from the idea that single stranded DNA or RNA molecules can be electrophoretically driven in a strict linear sequence through a biological pore that can be less than eight nanometers, and can be detected given that the molecules release an ionic current while moving through the pore. The pore contains a detection region capable of recognizing different bases, with each base generating various time specific signals corresponding to the sequence of bases as they cross the pore which are then evaluated.[104] Precise control over the DNA transport through the pore is crucial for success. Various enzymes such as exonucleases and polymerases have been used to moderate this process by positioning them near the pore's entrance.[105]
Short-read sequencing methods edit
Massively parallel signature sequencing (MPSS)edit
The first of the high-throughput sequencing technologies, massively parallel signature sequencing (or MPSS, also called next generation sequencing), was developed in the 1990s at Lynx Therapeutics, a company founded in 1992 by Sydney Brenner and Sam Eletr. MPSS was a bead-based method that used a complex approach of adapter ligation followed by adapter decoding, reading the sequence in increments of four nucleotides. This method made it susceptible to sequence-specific bias or loss of specific sequences. Because the technology was so complex, MPSS was only performed 'in-house' by Lynx Therapeutics and no DNA sequencing machines were sold to independent laboratories. Lynx Therapeutics merged with Solexa (later acquired by Illumina) in 2004, leading to the development of sequencing-by-synthesis, a simpler approach acquired from Manteia Predictive Medicine, which rendered MPSS obsolete. However, the essential properties of the MPSS output were typical of later high-throughput data types, including hundreds of thousands of short DNA sequences. In the case of MPSS, these were typically used for sequencing cDNA for measurements of gene expression levels.[59]
Polony sequencingedit
The polony sequencing method, developed in the laboratory of George M. Church at Harvard, was among the first high-throughput sequencing systems and was used to sequence a full E. coli genome in 2005.[106] It combined an in vitro paired-tag library with emulsion PCR, an automated microscope, and ligation-based sequencing chemistry to sequence an E. coli genome at an accuracy of >99.9999% and a cost approximately 1/9 that of Sanger sequencing.[106] The technology was licensed to Agencourt Biosciences, subsequently spun out into Agencourt Personal Genomics, and eventually incorporated into the Applied Biosystems SOLiD platform. Applied Biosystems was later acquired by Life Technologies, now part of Thermo Fisher Scientific.
454 pyrosequencingedit
A parallelized version of pyrosequencing was developed by 454 Life Sciences, which has since been acquired by Roche Diagnostics. The method amplifies DNA inside water droplets in an oil solution (emulsion PCR), with each droplet containing a single DNA template attached to a single primer-coated bead that then forms a clonal colony. The sequencing machine contains many picoliter-volume wells each containing a single bead and sequencing enzymes. Pyrosequencing uses luciferase to generate light for detection of the individual nucleotides added to the nascent DNA, and the combined data are used to generate sequence reads.[69] This technology provides intermediate read length and price per base compared to Sanger sequencing on one end and Solexa and SOLiD on the other.[78]
Illumina (Solexa) sequencingedit
Zdroj:https://en.wikipedia.org?pojem=DNA_sequencingText je dostupný za podmienok Creative Commons Attribution/Share-Alike License 3.0 Unported; prípadne za ďalších podmienok. Podrobnejšie informácie nájdete na stránke Podmienky použitia.
Antropológia
Aplikované vedy
Bibliometria
Dejiny vedy
Encyklopédie
Filozofia vedy
Forenzné vedy
Humanitné vedy
Knižničná veda
Kryogenika
Kryptológia
Kulturológia
Literárna veda
Medzidisciplinárne oblasti
Metódy kvantitatívnej analýzy
Metavedy
Metodika
Text je dostupný za podmienok Creative
Commons Attribution/Share-Alike License 3.0 Unported; prípadne za ďalších
podmienok.
Podrobnejšie informácie nájdete na stránke Podmienky
použitia.
www.astronomia.sk | www.biologia.sk | www.botanika.sk | www.dejiny.sk | www.economy.sk | www.elektrotechnika.sk | www.estetika.sk | www.farmakologia.sk | www.filozofia.sk | Fyzika | www.futurologia.sk | www.genetika.sk | www.chemia.sk | www.lingvistika.sk | www.politologia.sk | www.psychologia.sk | www.sexuologia.sk | www.sociologia.sk | www.veda.sk I www.zoologia.sk
